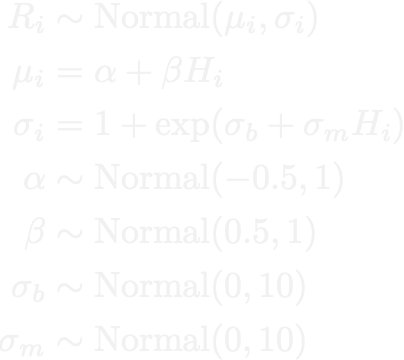Several times this year, I’ve had an idea for a post that never came to fruition. I figured a good way to wrap up the year would be to collect some of those loose threads into a single lightning-round collection of thoughts.
So, without further ado, here’s what I did in 2025:
I wrote a lot of R
A little over a year ago, I switched groups within our analytics team, and now have a strong requirement to use R for building projects. As a result, 2025 was the first year where I wrote more lines of R than any other language.
A few reflections after a year of working in R full-time:
-
I used to advise people who wanted to break into baseball analytics that it didn’t matter what language they learned, just pick R or Python and learn as much as you can. If a job requires you to switch, you can learn the other on the job, but prioritize knowledge depth while learning. This year has reinforced that opinion; good design patterns are ubiquitous, and learning on the fly has not been arduous. Living in a world with LLMs helps too (more on that later).
-
That being said, I’m now of the opinion that it’s near impossible to be a proper statistician and not engage with some R. There are several non-negotiable statistics libraries that plainly don’t have reasonable non-R alternatives (cough
mgcvcough). Certainly, there are ways around it if you go the Python route, but often it’s fitting a square peg in a round hole. -
I still prefer working in Python, especially for production code. R has so many idiosyncrasies that still frustrate me. The pain point that continues to plague me more than any is silent failures and tragically uninformative error messages. (Seriously, what does
object of type 'closure' is not subsettableeven mean?)
I made some small contributions to tidymodels
Speaking of R, after some initial skepticism, I’ve grown pretty fond of the tidymodels framework.
I'm here to walk back this take after 3 months. The upfront pain provides a lot of really good guardrails against doing really stupid statistical malpractice, and also makes downstream stuff (e.g. model tuning) trivially easy.
— Tyler Burch (@tylerjamesburch.com) March 7, 2025
Following my personal policy of helping out the libraries that help me, I made a couple of small PRs to tidymodels, probably the most interesting being adding fold weighting to the tune package.
For context, tune handles hyperparameter tuning within tidymodels. For each candidate hyperparameter set, you evaluate performance across resampling folds and take the set with the best average performance. Prior to this change, each fold contributed equally to that average, regardless of how much data it represented. That assumption is fine in a classic K-Fold setup, but if you switch to something like expanding window CV where folds have variable sizes, it seemed to miss the mark.
With this change, folds can now be weighted (naturally by training set size in the expanding window example) so later, more informative folds carry more influence when selecting hyperparameters. It’s a very small tweak, but it fixed a real issue I ran into on a project.
I’ve thought a lot about forecasting problems
Related to the above, more than before, I’ve found myself approaching problems through a forecasting lens. The largest project I worked on this year wasn’t explicitly a forecasting task, but required conditioning on time due to distributional shifts over time.
One of the clearest implications was cross-validation. I have increasingly used rolling or expanding window setups for cases when time could even plausibly matter, even if the problem isn’t explicitly framed as forecasting.
Lately, I’ve found myself defaulting to the assumption that time is relevant unless proven otherwise. Instead of asking whether I should use time-aware cross-validation, I enter with the posture that I need to be convinced it’s safe not to. Stationarity assumptions are helpful when true, but frequently break down in real-world problems, and ignoring that can lead to misleading performance estimates. Perhaps it can be a bit overcautious, but it’s one of those cases where a bit more care here has made me more comfortable with the results I’m delivering.
I used a lot of tokens
Agentic AI was impossible to ignore in 2025. I’ve long been bought in on codebase-aware LLMs. I find copy/pasting from ChatGPT both painfully slow and prone to stripping useful information. Because of this, I was a pretty early adopter of Cursor; I started using it in August of 2023. The tab complete was enough for me to buy in, and when agent-based edits came along, it became an important part of my workflow.
These days I’m using some combination of Cursor, Codex, and Claude Code for scaffolding boilerplate, generating prototypes (especially front-ends), quickly testing hypotheses, and making publication-quality plots faster than I could myself. The domain of tasks where it’s faster to prompt than to tweak, dissect, debug, etc. has grown way faster than I expected.
I don’t have any novel insights in this domain that haven’t been said elsewhere. The advice that I think about most day-to-day in my workflow is:
- Be meticulous about context. Keep context window usage under 50% if possible. The smaller the haystack, the easier the needle is to find.
- Spend time on prompting well. An extra 5 minutes on a good prompt can save an hour of debugging.
- Be judicious about correctness. One “wrong” line in the context window can yield hundreds of bad lines of code, or subtle unexpected bugs. Clear, correct prompts are key.
- Let the tool actually run code and see output. They’re better about this in other languages, but I find particularly in R they have to be pushed to do this. Add logging statements so it can iterate with itself and find bugs.
One thing I have noticed is that most of the public conversation around LLM tooling is focused more in the domain of traditional software engineering, where specs are far clearer than data analysis workflows. The projects I work on are fuzzier, and the road from question to answer (or from idea to predictions) is not typically well-defined from the outset. I haven’t seen nearly as much written about what works well in this environment and am still figuring it out day-by-day for myself.
I loosened my grip on statistical dogma
My first experience reading an honest stats text was McElreath’s Statistical Rethinking back in 2020 (10/10 recommend). Much of my early experience with doing statistics was strongly through a Bayesian lens. I used to feel pretty strongly that this was the best way to do things when possible.
In 2025, I let go of a lot of those biases. At the end of the day, I’m a practitioner; I need answers to problems. I’m not debating or writing about the philosophy of statistics, and in many of the problems I work on trying to wedge Bayesian inference into the solution can be a hindrance more than a value add.
If a frequentist framework can get me to effectively the same answer more quickly, I’ve become much more comfortable using it. In a lot of applied settings, the difference between a Bayesian posterior and a well-validated frequentist estimate is functionally negligible, while the difference in iteration speed isn’t. The Bayesian paradigm still shapes how I think about uncertainty, but I don’t reach for that machinery unless it provides something that will be used and I’m willing to wait for MCMC chains to finish sampling before I can provide an answer.
One of my repeated lines throughout this year has been “just use the right tool for the job,” which could even be a hidden thread behind this post. For me, the “right” tool is the one that answers the real question under real constraints and produces results that stakeholders can understand and act on. In practice, that’s often the approach that generates the most business value (or in my case, wins the most games) as quickly and correctly as possible.
I changed a lot of diapers
Above all else, I welcomed a second child into the world in May. At the time of writing this, I’m parenting a teething 7-month-old who gives the best smiles, as well as a curious and fiercely independent daughter who turned 3 yesterday.
I have a tendency to value myself entirely based on my work and the things I produce. Fatherhood is a constant forcing function to get out of that mindset and to enjoy life outside of sheer production. While it has exhausting moments, it has made me appreciate the day-to-day moments so much more.
Right now I’m most appreciating the sense of wonder from my toddler. I dread air travel, airports are awful, logistics are a nightmare, I could go on for hours. But hearing my daughter say “I’m so excited” while getting on a plane, and watching her stare with awe out the window during takeoff, has made me stop for a few moments and appreciate how cool life really can be. There are countless places where getting a chance to look through her lens has made me far more appreciative of the little things in life.

See you soon, hopefully before another 2 years go by, but no promises.
]]>